MOP-MCML 3D Extension: Full-Volume Optical Path Visualization on a Cartesian Grid
Following the baseline MOP-MCML and GPU-accelerated 2D version, we faced a fundamental limitation: a 2D cylindrical OP map cannot faithfully represent off-axis, asymmetric source-detector geometries. This post introduces the 3D extension of MOP-MCML, upgrading photon path recording from $(r, z)$ cylindrical to a full $(x, y, z)$ Cartesian grid, enabling complete volumetric visualization.
1. Why a 3D Version?
Standard MCML and earlier MOP-MCML accumulate photon weights in cylindrical coordinates $(r, z)$. This is efficient and accurate when the source is on-axis, but has inherent limitations:
- Off-axis source placement (non-rotationally symmetric geometries) causes cylindrical averaging to blur lateral asymmetry.
- X and Y lateral dimensions cannot be independently visualized.
- For multi-detector or asymmetric geometries (e.g., wrist-band sensors, ankle models), only a 3D view can truthfully reveal the photon penetration volume.
The 3D version introduces the OP_3D[nx × ny × nz] array to record full Cartesian optical path density, resolving all of the above.
2. Core Data Structure Changes
2.1 Input Structure (InputStruct)
On top of the existing dz, dr, nz, nr fields, four new fields are added for the X and Y grid dimensions:
1 | double dx; /* X grid spacing [cm] */ |
The .mci input file format is extended accordingly:
1 | 0.01 0.01 0.01 0.01 # dz, dr, dx, dy |
2.2 Output Structure (OutStruct)
A flat 1D array representing the 3D grid is added:
1 | double *OP_3D; /* 3D optical path density [nx * ny * nz] */ |
Indexing follows row-major (C-order): OP_3D[ix * ny * nz + iy * nz + iz], which can be directly reshaped into a 3D matrix in MATLAB or Python.
3. Photon Path Recording Mechanism
During each HopDropSpin call, four static buffers record the position and weight of every step of the current photon:
1 | static double sOut[60000] = {0}; /* accumulated weight at each step */ |
These contributions are only flushed into OP_3D after the photon is captured by a detector (hits the PD window). If the photon exits without being detected, the buffer is discarded without writing to OP_3D. This deferred-write mechanism ensures that OP_3D exclusively contains paths from photons that contribute to the detected signal, giving it a clear physical meaning.
4. PD_MODE Compile-Time Option
The 3D version selects the detector mode via a compile-time macro, eliminating runtime branching overhead:
PD_MODE |
Meaning |
|---|---|
1 |
Count only the reflection detector (top surface, $z \approx 0$) |
2 |
Count only the transmission detector (bottom surface, $z \approx z_{max}$) |
3 |
Count both reflection and transmission (default) |
Compilation example (reflection-only mode):
1 | gcc -O3 -DPD_MODE=1 -o local_mcml_3d mcmlmain.c mcmlgo.c mcmlio.c mcmlnr.c -lm |
The batch script run_3d_variants.sh automatically compiles, runs, and renames outputs for all three modes:
1 | bash run_3d_variants.sh |
After completion, the working directory contains:
1 | 3d_cpu_Ronly.mco |
5. Input File Example
A typical 3D simulation input file test_3d.mci:
1 | 1.0 # file version |
Key parameters:
- nx=300, ny=300, nz=200: Total voxels = $300 \times 300 \times 200 = 18 \times 10^6$; stored as
doublerequires ~144 MB RAM, well within normal range. - PD windows: Reflection PD centered at $(0.2, 0)$ cm, side length $0.1$ cm; Transmission PD centered at $(-0.2, 0)$ cm, side length $0.1$ cm.
- Light source: Point source at $(-0.2, 0)$ cm, co-axial with the transmission detector.
6. 3D Visualization
We developed the MATLAB script look_mop_3d.m to interactively explore the OP_3D data through draggable cross-sectional planes.
6.1 Interactive Slice Viewer
The script simultaneously renders three orthogonal semi-transparent slices (X-plane, Y-plane, Z-plane) in the 3D volume, color-coded as $\log_{10}(\text{OP})$ in units of $1/\text{cm}^3$. Three sliders at the bottom of the figure control slice positions in real time — dragging any slider instantly updates the corresponding plane without recomputation:
- X slider: Moves the YZ slice — observe depth-wise photon distribution at different lateral X positions.
- Y slider: Moves the XZ slice — compare scattering on the source side vs. the detector side.
- Z slider: Moves the XY slice — inspect the lateral photon spot size at a specific depth (e.g., skin layer, muscle layer).
6.2 Usage
In MATLAB, edit the parameters at the top of the script, then run:
1 | mco_file = 'validate_RT/3d_cpu_RT.mco'; % target .mco file |
1 | run('look_mop_3d.m') |
6.3 Simulation Result
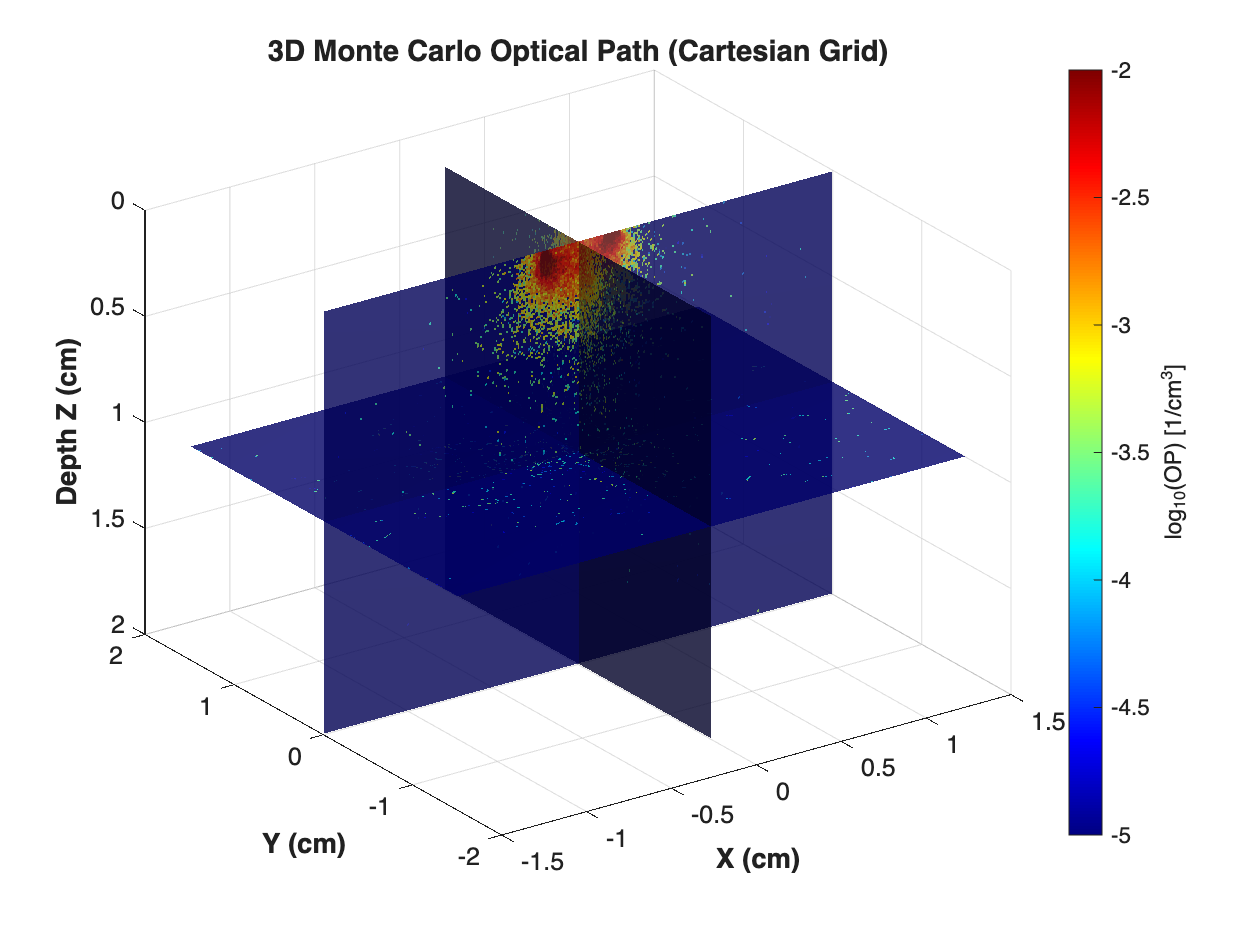
The figure below shows the actual output of the CPU simulation: three orthogonal cross-sectional slices of the $\log_{10}(\text{OP})$ volumetric field at initial positions (X=0, Y=0, Z=mid-depth). The photon injection point (point source at the top surface $z=0$) has the highest density (dark red), decaying with depth and lateral distance. This is the authoritative output of look_mop_3d.m applied to 3d_cpu_RT.mco.
6.4 Web Interactive Explorer
Note: The interactive viewer below is built for exploration on this webpage. It uses a spatially downsampled version of the same dataset (factor ×2, i.e., $150 \times 150 \times 100$ voxels vs. the original $300 \times 300 \times 200$). The color scale and photon distribution pattern are faithful representations of the data, but the exact values and fine-grained spatial resolution should be taken from the authoritative figure above.
How to use:
- Drag the 3D figure (left-click + drag) to rotate the view.
- Use the sliders or ± buttons to move each cross-sectional plane along X, Y, or Z. You can also type a number directly into the input box.
- The X slider moves the YZ plane laterally (observe depth-wise asymmetry).
- The Y slider moves the XZ plane (compare source vs. detector side).
- The Z slider moves the XY horizontal plane (inspect photon spot size at a given depth layer).
7. GPU 3D Version
The 3D optical path computation also has a corresponding GPU (CUDA) implementation, sharing the same physical model (Hop-Drop-Spin) and OP_3D output format as the CPU version. The two implementations produce statistically equivalent results — the only difference is computation speed.
7.1 Accuracy Equivalence Validation
The validate_RT directory contains CPU and GPU outputs for all three PD modes, ready for comparison:
| File | Version | Mode |
|---|---|---|
3d_cpu_RT.mco |
CPU | RT |
3d_gpu_RT.mco |
GPU | RT |
3d_cpu_Ronly.mco |
CPU | R-only |
3d_gpu_Ronly.mco |
GPU | R-only |
3d_cpu_Tonly.mco |
CPU | T-only |
3d_gpu_Tonly.mco |
GPU | T-only |
Comparing Rd and Tt values (from summary.csv) and the OP_3D slice plots confirms that both versions agree within statistical fluctuations, validating the physical correctness of the GPU parallel implementation.
7.2 Performance Advantage
The 3D grid upgrades OP_3D from the earlier 2D array to a full $N_x \times N_y \times N_z$ volume. The substantially higher memory access and computation load makes GPU acceleration especially advantageous here:
| Metric | CPU (local) | GPU (A100) |
|---|---|---|
| Photons | $10^6$ | $10^6$ |
| Grid size | $300\times300\times200$ | $300\times300\times200$ |
| Runtime | ~minutes | < 1 s |
| Output accuracy | Baseline | Statistically equivalent |
7.3 Running the GPU 3D Version
The GPU 3D version lives in the dedicated version_3d_gpu/ directory (separate from the 2D GPU code). Submit via SLURM on the LIP6 Convergence cluster (A100 partition):
1 | cd version_3d_gpu |
The script automatically compiles with nvcc and outputs 3d_gpu_Ronly_A.mco, 3d_gpu_Tonly_A.mco, and 3d_gpu_RT_A.mco. To compile manually without SLURM:
1 | nvcc -O3 -arch=sm_80 -DPD_MODE=3 -o mcml_3d_gpu mcml_gpu.cu mcmlio_gpu.c |
Once done, load any GPU output directly into look_mop_3d.m for interactive visualization:
1 | mco_file = 'validate_RT/3d_gpu_RT.mco'; |
8. Performance and Limitations
| Metric | CPU (local, Apple M-series) | GPU (A100) |
|---|---|---|
| Photons | $10^6$ | $10^6$ |
| Grid size | $300\times300\times200$ | $300\times300\times200$ |
| Memory (OP_3D) | ~144 MB (RAM) | ~144 MB (VRAM) |
| Runtime | ~minutes | < 1 s |
| Result accuracy | Yes | Yes (equivalent) |
Key limitations:
- 3D grid memory scales cubically with resolution ($N_x \times N_y \times N_z$); high-resolution scenarios should use the GPU version to keep runtimes manageable.
- The static buffer
sOut[60000]caps the maximum number of steps per photon; photons with very long paths (e.g., in near-zero scattering media) may hit this boundary.