MOP-MCML GPU Version: 175x Faster 2D Light Transport Simulation
Project Background
The Monte Carlo Multi-Layered (MCML) method is the gold standard algorithm in biomedical optics for simulating photon transport in multi-layered turbid media. By statistically tracing large numbers of random-walking photons, it estimates key physical quantities — reflectance ($R_d$), transmittance ($T_t$), and the Mean Optical Path (MOP, $L_{eff}$) — providing a theoretical basis for optical diagnosis, photoplethysmography (PPG), and photodynamic therapy.
This project implements both a CPU and a high-performance GPU version of the 2D MOP-MCML simulator. The original CPU implementation was developed by Songlin Li. It was extended with GPU acceleration and improved I/O by Wenzheng Wang under the supervision of Prof. Sylvain Feruglio at the LIP6 Laboratory, Sorbonne University.
Note: For the 3D Cartesian grid extension (full-volume OP visualization), see MOP-MCML 3D Version.
Why GPU Acceleration?
The CPU version processes photons serially. While sufficient for hundreds of thousands of photons, computation time grows prohibitively for the millions of photons required for high statistical accuracy. Simulating 2 million photons on a CPU takes approximately 70 seconds per wavelength. A full parameter scan across tissue thicknesses and wavelengths can therefore take hours.
The GPU’s massively parallel architecture is a natural fit: each photon’s transport path is statistically independent and follows identical computational logic — a textbook case for data-level parallelism.
Core Features of the GPU Version
1. Significant Performance Improvement
Benchmarked on an NVIDIA A100 GPU:
| Metric | CPU Version | GPU Version (CUDA) | Speedup |
|---|---|---|---|
| Runtime | ~70.0 s | ~0.4 s | 175× |
| Photon execution | Serial | Thousands in parallel | — |
| Precision | Double | Double + atomic ops | — |
Test condition: 2 million photons per run on a single layer tissue model.
A full scan that previously required hours now completes in minutes. For iterative inverse-problem workflows or multi-wavelength studies, this speedup is decisive.
2. Implementation Highlights
The GPU implementation is a deep re-engineering for the parallel computing paradigm, not a naïve code port:
- Parallel RNG: Uses the CUDA
curandlibrary to generate independent high-quality random number streams per thread, eliminating correlation artifacts. - Atomic accumulation: Photon weight contributions to
Rd_ra,A_rz, andTt_raarrays useatomicAddto guarantee correctness under concurrent writes from thousands of threads. - Flattened memory layout: Multi-dimensional output arrays are stored in contiguous 1D blocks, maximizing coalesced global memory access on the GPU.
- Double precision throughout: The full double-precision arithmetic of the CPU version is preserved, ensuring numerical equivalence.
The physical Hop-Drop-Spin algorithm remains identical to the CPU version; only the execution model changes from serial to massively parallel.
3. Batch Workflow and Auto-CSV Summary
A key practical addition is automatic per-run CSV summary generation. After each simulation, the program appends a row to a summary_<label>.csv file:
1 | output, Rd, Tt |
This makes it easy to aggregate results from a parameter sweep without post-processing scripts.
Local Compilation and Execution
On a workstation with an NVIDIA GPU:
1 | cd version_gpu |
HPC Cluster Integration (SLURM)
For large-scale parameter scanning on the LIP6 Convergence cluster (A100 partition):
1 | # Single run |
The run_batch.slurm script recompiles the code, removes stale .mco files, and sequentially runs 3mm_1500.mci, 4mm_1500.mci, and 5mm_1500.mci:
1 |
|
For multi-mode variant runs (Ronly / Tonly / RT), use run_gpu_variants.slurm — this also serves as the entry point for the 3D GPU extension.
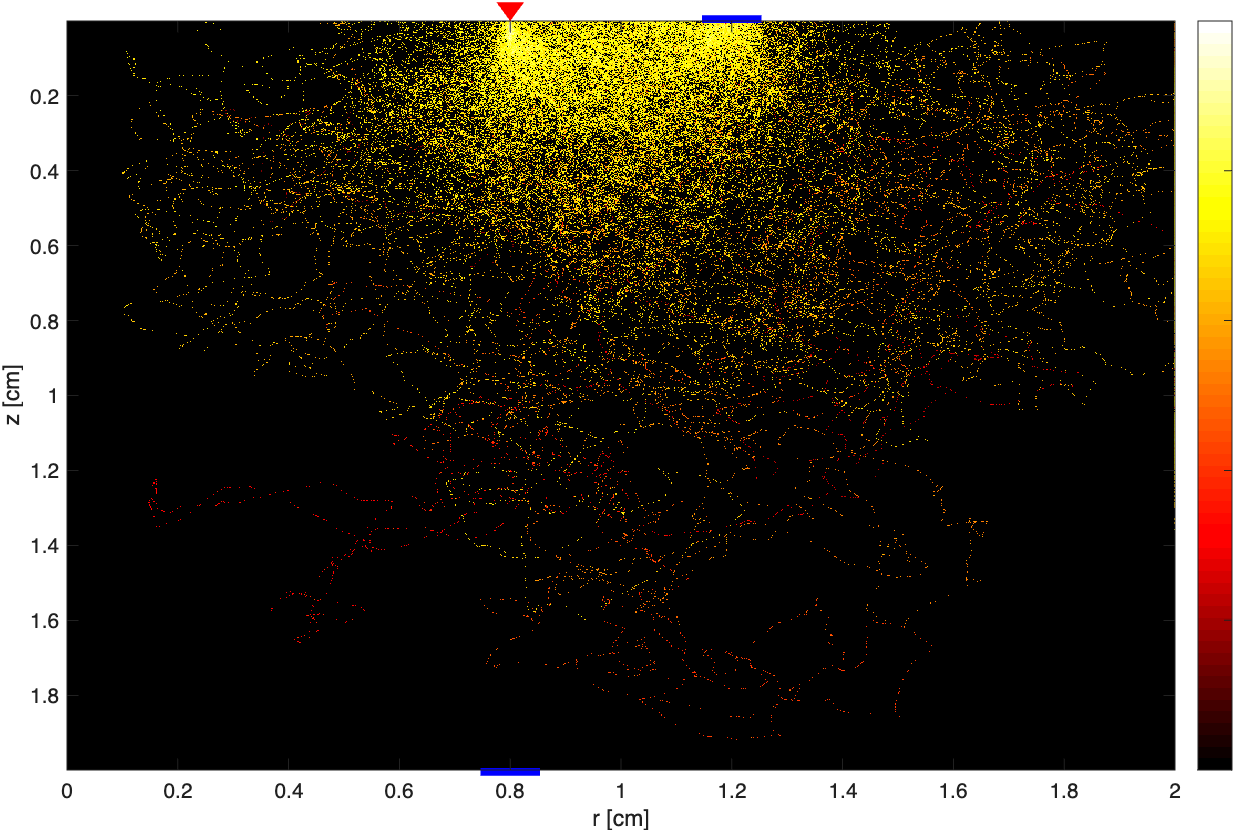
4. Visualization Tools
Beyond MATLAB scripts, a Python visualization tool plot_results.py generates publication-ready figures (300 DPI):
- Reflectance ($R_d$) curves as a function of wavelength, for 3 mm / 4 mm / 5 mm tissue.
- Transmittance ($T_t$) trends across wavelengths and thicknesses.
- Multi-condition overlay plots for comparing the influence of optical parameters.
1 | python3 version_gpu/plot_results.py |
Application Scenarios
The GPU-accelerated 2D version is particularly well-suited for:
- Large-scale parameter sweeps: Rapidly scanning wavelength, tissue thickness, and $\mu_a / \mu_s$ combinations.
- Inverse problem solving: Iteratively retrieving tissue optical properties from measured $R_d$ / $T_t$ data.
- Sensor design optimization: Finding the optimal source-detector separation (SDS) for wearable or implantable photonic sensors.
- Educational demonstrations: Near-instant classroom demos of how tissue properties alter light transport.
Technical Requirements
- Hardware: NVIDIA GPU (Compute Capability 5.0+; A100 / V100 / RTX A-series recommended for research).
- Software: CUDA Toolkit ≥ 11.8, GCC ≥ 9.
For users without a GPU, the CPU version remains available for small-scale verification and debugging.
Summary
The GPU port of MOP-MCML reduces 2D simulation time by two orders of magnitude — from ~70 s to ~0.4 s per run. Combined with automatic CSV logging, HPC SLURM integration, and the Python visualization pipeline, it forms a complete parameter-scanning workflow for biomedical optics research.
For scenarios requiring full volumetric 3D resolution, see the MOP-MCML 3D Extension, which extends the same GPU engine to a Cartesian $N_x \times N_y \times N_z$ grid with interactive MATLAB slice viewer.
Project Link: GitHub - Pasdeau/MOP-MCML